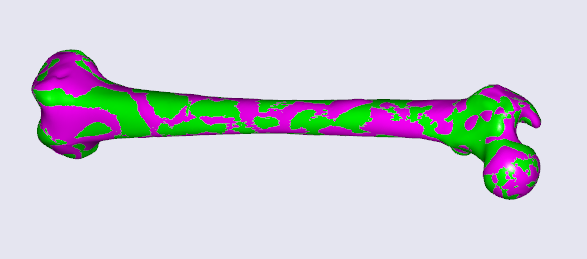
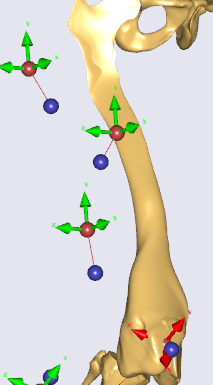
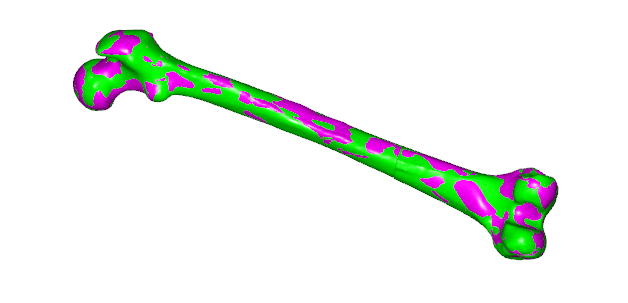
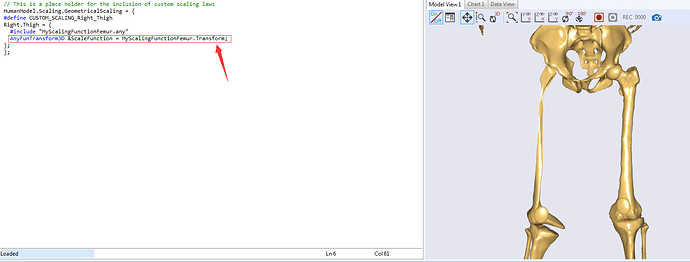
/* This file defines a subject-specific morphing law for the Lesson 4 of Scaling tutorial*/
AnyFolder MyScalingFunctionFemur = {
AnyFunTransform3DLin Transform = {
ScaleMat = {{1,0,0},{0,1,0},{0,0,1}};
Offset = {0,0,0};
PreTransforms = {&.STLTransform, &.ReverseTransform};
};
AnyFunTransform3DLin2 AffineTransform =
{
//PreTransforms = {};
Points0 = …TSeg2ScaleFrame(
{
{0.0776166,-0.38519,0.0434183},
{0.0291691,-0.388362,0.0489085},
{0.0702968,-0.375534,-0.0240044},
{0.0171157,-0.195817,0.013274},
{-0.0481888,-0.00649841,0.00942058},
{0.009532983,0.016264,-0.015843}
});
Points1 =
{
{-0.0335496,0.00561388,-0.0159026},
{0.0131622,0.0129517,-0.0401973},
{0.00955101,0.0160026,0.0352691},
{0.0253999,0.200918,-0.0145758},
{0.0797432,0.417332,-0.022623},
{0.0356885,0.431939,0.0166936}
};
Mode = VTK_LANDMARK_AFFINE;
};
AnyFunTransform3DLin2 ReverseTransform = {
Points0 = .AffineTransform.Points1;
Points1 = .AffineTransform.Points0;
Mode = VTK_LANDMARK_RIGIDBODY;
};
AnyFunTransform3DRBF RBFTransform =
{
PreTransforms = {&.AffineTransform};
RBFDef =
{
Type = RBF_ThinPlate;
Param = 1;
};
Points0 = …TSeg2ScaleFrame(
{
{0.0776166,-0.38519,0.0434183},
{0.0874067,-0.392876,0.0191814},
{0.0291691,-0.388362,0.0489085},
{0.0336871,-0.401527,0.0528365},
{0.0702968,-0.375534,-0.0240044},
{0.0840476,-0.405399,-0.0240044},
{0.046656,-0.36353,0.00203508},
{0.0519064,-0.422182,0.045868},
{0.6825022,-0.426252,-0.0176293},
{0.0171157,-0.195817,0.013274},
{-0.0153646,-0.0538598,-0.00275116},
{-0.0111436,-0.0677822,-0.000322446},
{-0.0481888,-0.00649841,0.00942058},
{0.009532983,0.016264,-0.015843},
{-0.0060929,-0.0615546,0.00151425},
{-0.036961,-0.0406715,0.053172},
{-0.038943,-0.0406992,0.0101967},
{0.0005855,-0.0509986,0.0225059},
{0.0885529,-0.388669,0.00365604},
{0.0606754,-0.420174,-0.0318583},
{0.055317,-0.422778,-0.00984486},
{0.0349846,-0.423029,0.0426269},
{0.052857,-0.420594,0.0475201},
{0.080049,-0.370333,0.0108282},
{0.0708826,-0.371049,0.0418323},
{0.0353599,-0.347786,0.0276989},
{0.0536242,-0.343088,0.00172468},
{0.0827935,-0.39489,-0.0240079},
{0.0169421,-0.390191,0.0277442},
{0.0162501,-0.410556,0.0348857},
{0.0284323,-0.386989,0.0139395},
{0.0177057,-0.398845,0.0187154},
{0.0389646,-0.396279,-0.025363},
{0.0888908,-0.401907,-0.00876824},
{0.0548423,-0.397694,0.054172},
{0.077992,-0.421114,0.00714761},
{0.0463893,-0.284064,0.011615},
{0.0632637,-0.3013,0.0116601},
{0.0558027,-0.254465,0.0189366},
{0.0487975,-0.316967,0.00655951},
{0.0640665,-0.335391,0.00255428},
{0.0535251,-0.272982,0.0154517},
{0.0405198,-0.22445,0.0146546},
{0.0213611,-0.169527,0.0158975},
{0.0377727,-0.194215,0.0374407},
{0.0502173,-0.234592,0.0301236},
{0.0506393,-0.239614,0.0207116},
{0.0231513,-0.417397,0.0260686},
{0.0774498,-0.38116,0.0283361},
{0.0749767,-0.375019,0.0308108}
});
Points1 = {
{-0.0335496,0.00561388,-0.0159026},
{-0.0280022,-0.00269058,0.0120236},
{0.0131622,0.0129517,-0.0401973},
{0.012141,-0.000433684,-0.0446263},
{0.00955101,0.0160026,0.0352691},
{0.00075525,-0.0115942,0.0405223},
{0.0119686,0.0280065,0.0120921},
{-0.00935188,-0.0231205,-0.0279828},
{-0.00432023,-0.02776477,0.0308913},
{0.0253999,0.200918,-0.0145758},
{0.0585843,0.361284,0.000857341},
{0.0550661,0.346257,-0.00206488},
{0.0797432,0.417332,-0.022623},
{0.0356885,0.431939,0.0166936},
{0.0503574,0.351978,0.00110147},
{0.06372,0.381583,-0.0571084},
{0.0809548,0.383419,-0.0216873},
{0.0338266,0.35835,-0.0121963},
{-0.024259,0.00394552,0.0217401},
{0.0194315,-0.0242202,0.0356052},
{0.0178418,-0.0246098,0.0145176},
{0.00503235,-0.0230989,-0.0367033},
{-0.0103818,-0.0219988,-0.0288968},
{-0.0198254,0.0267586,0.01261},
{-0.0264373,0.024819,-0.0170189},
{0.0114769,0.0436825,-0.0237335},
{0.00778656,0.0454752,0.00969556},
{0.00118152,0.00134793,0.0426428},
{0.0293845,0.00573355,-0.0302668},
{0.0303841,-0.00849202,-0.0380678},
{0.024226,0.00585466,-0.00989906},
{0.0320057,-0.0066778,-0.0235614},
{0.0327069,-0.00420313,0.0239168},
{-0.0118756,-0.0072693,0.0340803},
{-0.013875,0.00182269,-0.0345471},
{-0.0159371,-0.0209117,0.0101429},
{0.00910361,0.107057,-0.000237901},
{-0.0108107,0.083166,0.00767706},
{-0.00792776,0.133574,-0.00168584},
{0.00402107,0.0750955,0.00614671},
{-0.00431338,0.0517168,0.0142384},
{-0.00429717,0.115213,0.0035777},
{0.00607966,0.182325,-0.00565917},
{0.0207678,0.241608,-0.0108694},
{-0.00150775,0.198673,-0.0232213},
{-0.00827493,0.154543,-0.0107971},
{-0.0024134,0.158461,-0.00343703},
{0.0208062,-0.0232439,-0.0270419},
{-0.0251027,0.00861673,-0.00121048},
{-0.0240169,0.0216748,-0.00427663}
};
BoundingBox =
{
Type = BB_Cartesian;
ScaleXYZ = {2, 2, 2};
DivisionFactorXYZ = 5*{1, 1, 1};
};
BoundingBoxOnOff = On;
};
AnyFunTransform3DSTL STLTransform =
{
PreTransforms = {&.RBFTransform};
RBFDef.Type = RBF_ThinPlate;
AnyFixedRefFrame Input = {
AnySurfSTL SourceSurf = {
FileName = “femur211.stl”;
ScaleXYZ = {1, 1, 1};
};
AnySurfSTL TargetSurf = {
FileName = “TargetFemur.stl”;
ScaleXYZ = {1, 1, 1};
AnyFunTransform3D &pre = …TSeg2ScaleFrame;
};
};
SurfaceObjects0 = {&Input.SourceSurf};
SurfaceObjects1 = {&Input.TargetSurf};
NumPoints = 400;
BoundingBox.ScaleXYZ = {2, 2, 2};
BoundingBox.DivisionFactorXYZ = {1, 1, 1};
BoundingBoxOnOff = On;
};
}; // MyScalingFunctionFemur
 . However, I gave up because I do not know what I’m doing wrong. That’s why I am asking for help, how should I personalize my model based on image data.
. However, I gave up because I do not know what I’m doing wrong. That’s why I am asking for help, how should I personalize my model based on image data.